McUtils.Zachary
Handles much of the higher-order “numerical math” stuff inside Mcutils which has made it balloon a little bit Deals with anything tensor, Taylor expansion, or interpolation related
Members
Examples
1D finite difference derivative via finite_difference:
from McUtils.Zachary import finite_difference
import numpy as np
sin_grid = np.linspace(0, 2*np.pi, 200)
sin_vals = np.sin(sin_grid)
deriv = finite_difference(sin_grid, sin_vals, 3) # 3rd deriv
base = Plot(sin_grid, deriv, aspect_ratio = .6, image_size=500)
Plot(sin_grid, np.sin(sin_grid), figure=base)
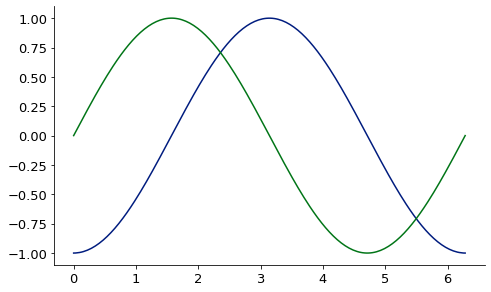
2D finite difference derivative via finite_difference:
from McUtils.Zachary import finite_difference
import numpy as np
x_grid = np.linspace(0, 2*np.pi, 200)
y_grid = np.linspace(0, 2*np.pi, 100)
sin_x_vals = np.sin(x_grid); sin_y_vals = np.sin(y_grid)
vals_2D = np.outer(sin_x_vals, sin_y_vals)
grid_2D = np.array(np.meshgrid(x_grid, y_grid)).T
deriv = finite_difference(grid_2D, vals_2D, (1, 3))
TensorPlot(np.array([vals_2D, deriv]), image_size=500)
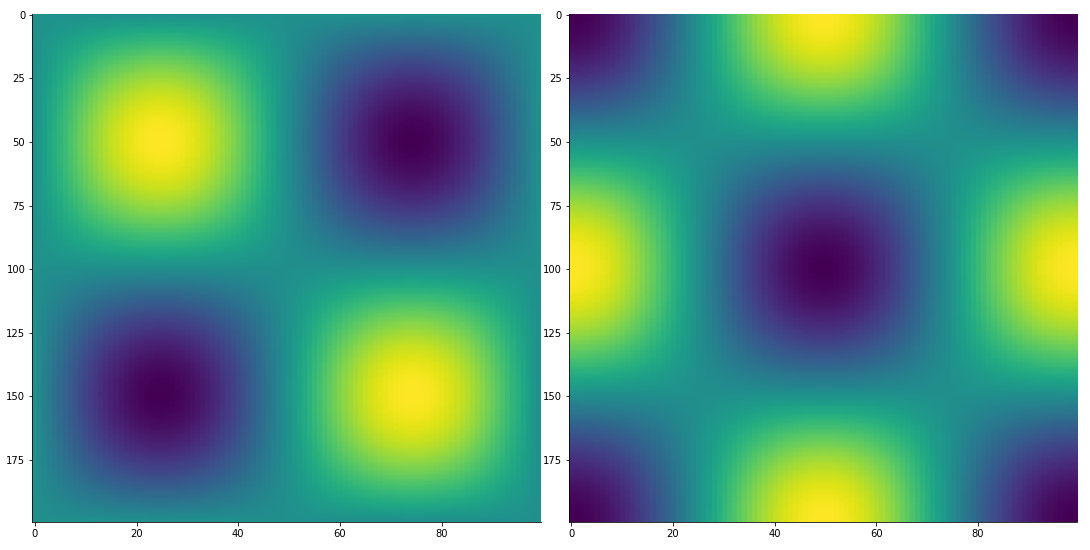
Create a convenient, low-order expansion of a (potentially expensive) function
def sin_xy(pt):
ax = -1 if pt.ndim>1 else 0
return np.prod(np.sin(pt), axis=ax)
point = np.array([.5, .5])
# create the function expansions
exp1 = FunctionExpansion.expand_function(sin_xy, point, function_shape=((2,), 0), order=1, stencil=5)
exp2 = FunctionExpansion.expand_function(sin_xy, point, function_shape=((2,), 0), order=2, stencil=6)
exp4 = FunctionExpansion.expand_function(sin_xy, point, function_shape=((2,), 0), order=4, stencil=6)
# create a test grid and plot the approximations
test_grid = np.vstack([np.linspace(-.5, .5, 100), np.zeros((100,))]).T + point[np.newaxis]
g = test_grid[:, 0]
gg = GraphicsGrid(nrows=1, ncols=3, subimage_size=350)
for i, e in zip(range(3), (exp1, exp2, exp4)):
# plot the real answer
gg[0, i] = Plot(g, sin_xy(test_grid), figure=gg[0, i])
# plot the expansion
gg[0, i] = Plot(g, e(test_grid), figure=gg[0, i])
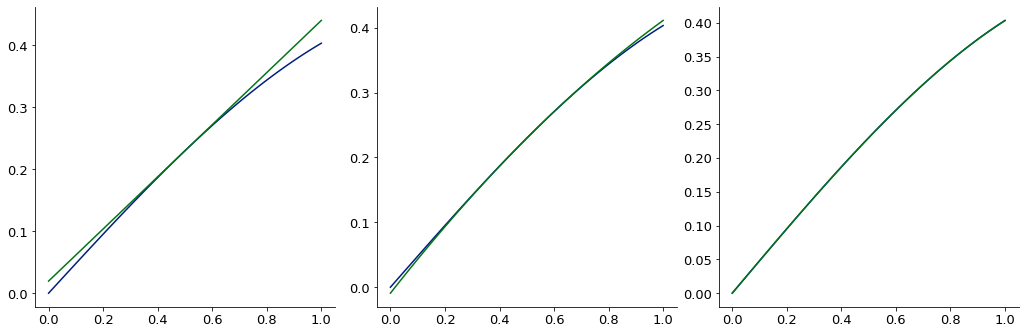
expansions work in multiple dimensions, too
mesh = np.meshgrid(np.linspace(.4, .6, 100, dtype='float128'), np.linspace(.4, .6, 100, dtype='float128'))
grid = np.array(mesh).T
gg2 = GraphicsGrid(nrows=2, ncols=3, subimage_size=350)
# plot error in linear expansion
styles = dict(ticks_style=(False, False), plot_style={'vmin': np.min(sin_xy(grid)), 'vmax': np.max(sin_xy(grid))})
gg2[0, 0] = ContourPlot(*mesh, sin_xy(grid), figure=gg2[0, 0], **styles)
gg2[0, 1] = ContourPlot(*mesh, exp1(grid), figure=gg2[0, 1], **styles)
# when we plot the error, we shift it so that it's centered around the average function value to show the scale
# of the error
gg2[0, 2] = ContourPlot(*mesh,
np.marginalize_out([np.min(sin_xy(grid)), np.max(sin_xy(grid))]) + sin_xy(grid) - exp1(grid),
figure=gg2[0, 2], **styles)
# plot error in quadratic expansion
gg2[1, 0] = ContourPlot(*mesh, sin_xy(grid), figure=gg2[1, 0], **styles)
gg2[1, 1] = ContourPlot(*mesh, exp2(grid), figure=gg2[1, 1], **styles)
gg2[1, 2] = ContourPlot(*mesh,
np.marginalize_out([np.min(sin_xy(grid)), np.max(sin_xy(grid))]) + sin_xy(grid) - exp2(grid),
figure=gg2[1, 2], **styles)
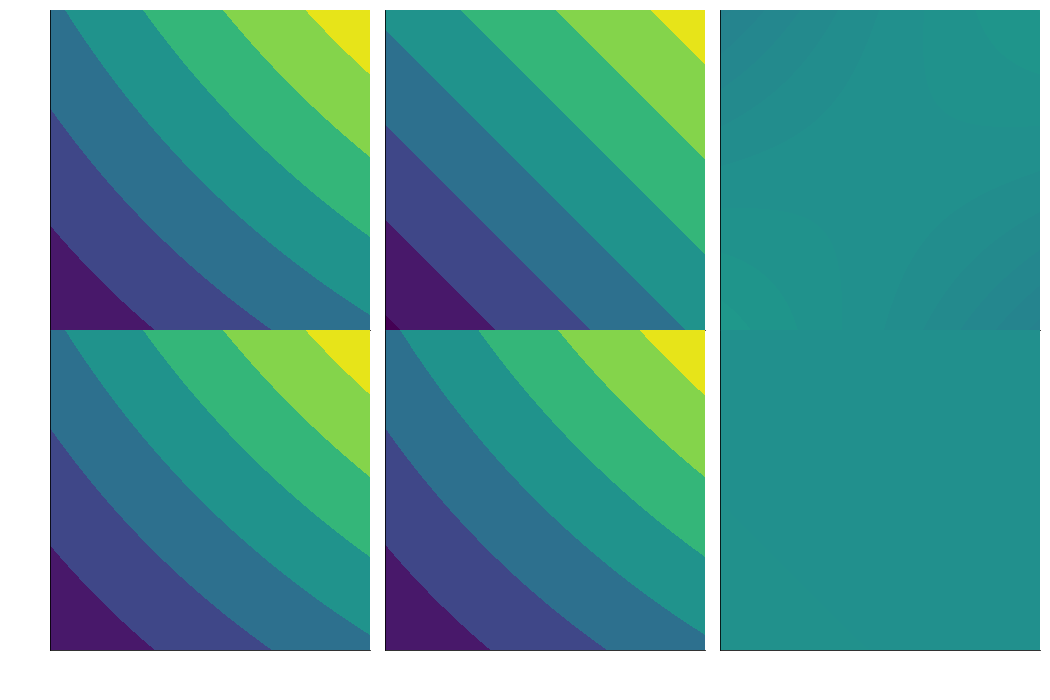

- stirs
- bin_gs
- bins
- facs
- fd_weights
- uneven_weights
- finite_difference
- FD2D
- FD2D_multi
- FDDeriv
- FDDeriv2
- FiniteDifferenceParallelism
- TensorTerms
- TensorConversion
- PseudopotentialTerms
- ExplicitPoly
- TensorDerivatives
- ExpandFunction
- ExpansionDerivs
- MultiExpansion
- Polys
- SparseShifts
- SparseArrayPoly
- SparseConvolve
- LinSpaceMesh
- MeshGridMesh
- StructuredMesh
- UnstructuredMesh
- RegularMesh
- RegMeshSubgrids
- SemiStructuredMesh
- MeshFromList
- ParametricCurveInterpolation
- RegularGridInterpolatorND
- Interpolator1D
- InterpolatorExtrapolator2D
- InterpolatorExtrapolator1D
- RegularInterpolator3D
- IrregularInterpolator3D
- TensorExpressionEfficiency
- RBFInterp1D
- RBFInterpolator
- RBFInterpolatorResiliance
- HigherElementaryDerivs
- RBFTiming
- RBFForms
- Symbolics
- CoordinateFunctions
- InternalReexpansions
Before we can run our examples we should get a bit of setup out of the way. Since these examples were harvested from the unit tests not all pieces will be necessary for all situations.
All tests are wrapped in a test class
class ZacharyTests(TestCase):
def setUp(self):
self.save_data = TestManager.data_gen_tests
def __getstate__(self):
return {}
def __setstate__(self, state):
pass
def get_error(self, ref_vals, vals):
err = np.abs(vals - ref_vals)
cum_err = np.linalg.norm(err.flatten())
max_err = np.max(err.flatten())
mean_err = np.average(err.flatten())
return err, cum_err, max_err, mean_err
def plot_err(self, grid, ref_vals, vals, errs):
if grid.ndim == 1:
Plot(grid, ref_vals, figure=Plot(grid, vals))
Plot(grid, errs[0]).show()
elif grid.ndim == 3:
g = (grid[:, :, 0], grid[:, :, 1])
gg = GraphicsGrid(nrows=2, ncols=2)
gg[0, 0] = ContourPlot(*g, ref_vals, figure=gg[0, 0])
gg[1, 0] = ContourPlot(*g, vals, figure=gg[1, 0])
gg[1, 1] = ContourPlot(*g, errs[0], figure=gg[1, 1])
gg[0, 0].tight_layout()
gg.show()
def print_error(self, n, order, errs):
print(
"Order: {}.{}\nCumulative Error: {}\nMax Error: {}\nMean Error: {}".format(n, order, *errs[1:])
)
class harmonically_coupled_morse:
# mass_weights = masses[:2] / np.sum(masses[:2])
def __init__(self,
De_1, a_1, re_1,
De_2, a_2, re_2,
kb, b_e
):
self.De_1 = De_1
self.a_1 = a_1
self.re_1 = re_1
self.De_2 = De_2
self.a_2 = a_2
self.re_2 = re_2
self.kb = kb
self.b_e = b_e
def __call__(self, carts):
v1 = carts[..., 1, :] - carts[..., 0, :]
v2 = carts[..., 2, :] - carts[..., 0, :]
r1 = nput.vec_norms(v1) - self.re_1
r2 = nput.vec_norms(v2) - self.re_2
bend, _ = nput.vec_angles(v1, v2)
bend = bend - self.b_e
return (
self.De_1 * (1 - np.exp(-self.a_1 * r1)) ** 2
+ self.De_2 * (1 - np.exp(-self.a_2 * r2)) ** 2
+ self.kb * bend**2
)
def constructSparseArray(cls, shape, n_els):
from McUtils.Numputils import SparseArray
np.random.seed(1)
inds = np.unique(np.array([np.random.choice(x, n_els) for x in shape]).T, axis=0)
vals = np.random.rand(len(inds))
inds = inds.T
# `from_data` for backend flexibility
return SparseArray.from_data(
(
vals,
inds
),
shape=shape
)
stirs
def test_stirs(self):
stir = StirlingS1(8)
ans = np.array([
[1, 0, 0, 0, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0, 0, 0],
[0, -1, 1, 0, 0, 0, 0, 0],
[0, 2, -3, 1, 0, 0, 0, 0],
[0, -6, 11, -6, 1, 0, 0, 0],
[0, 24, -50, 35, -10, 1, 0, 0],
[0, -120, 274, -225, 85, -15, 1, 0],
[0, 720, -1764, 1624, -735, 175, -21, 1]
])
# print(stir, file=sys.stderr)
self.assertAlmostEqual(np.round(np.linalg.norm(stir-ans), 6), 0.)
bin_gs
def test_bin_gs(self):
gbin = GammaBinomial(7/2, 8)
ans = np.array([1., 3.5, 4.375, 2.1875, 0.273438, -0.0273438, 0.00683594, -0.00244141])
# print(gbin, file=sys.stderr)
self.assertAlmostEqual(np.round(np.linalg.norm(gbin-ans), 5), 0.)
bins
def test_bins(self):
gbin = Binomial(8)
ans = np.array([
[1, 0, 0, 0, 0, 0, 0, 0],
[1, 1, 0, 0, 0, 0, 0, 0],
[1, 2, 1, 0, 0, 0, 0, 0],
[1, 3, 3, 1, 0, 0, 0, 0],
[1, 4, 6, 4, 1, 0, 0, 0],
[1, 5, 10, 10, 5, 1, 0, 0],
[1, 6, 15, 20, 15, 6, 1, 0],
[1, 7, 21, 35, 35, 21, 7, 1]
])
# print(gbin, file=sys.stderr)
self.assertAlmostEqual(np.sum(gbin-ans), 0.)
facs
def test_facs(self):
facs = Factorial(8)
ans = np.array([1, 1, 2, 6, 24, 120, 720, 5040])
# print(facs, file=sys.stderr)
self.assertAlmostEqual(np.linalg.norm(facs-ans), 0.)
fd_weights
def test_fd_weights(self):
coeffs = RegularGridFiniteDifference.get_weights(3, 7/2, 7)
ans = [ -0.0192708, 0.259896, -2.02969, 4.92448, -4.92448, 2.02969, -0.259896, 0.0192708 ]
coeffs2 = RegularGridFiniteDifference.get_weights(3, 0, 7)
ans2 = [
-8.058333333333334, 42.53333333333333, -98.225,
129.66666666666666, -106.04166666666667, 53.6,
-15.408333333333333, 1.9333333333333333
]
# print(coeffs2, file=sys.stderr)
r1 = np.round(np.linalg.norm(coeffs-ans), 5)
r2 = np.round(np.linalg.norm(coeffs2-ans2), 10)
self.assertAlmostEqual(r1 + r2, 0.)
uneven_weights
def test_uneven_weights(self):
import numpy as np
weights = IrregularGridFiniteDifference.get_weights
uweights = [
weights(2, 0, np.array([-2, -1, 0, 1, 2])),
weights(1, 0, np.array([-3/2, -1/2, 1/2, 3/2])),
weights(1, 1, np.array([-3, -2, -1, 0, 1]))
#weights(3, 1/2, np.array([0, 1/3, 1, 2, 7/2, 6]))
]
eweights = RegularGridFiniteDifference.get_weights
targ_weights = [
eweights(2, 2, 4),
eweights(1, 3/2, 3),
eweights(1, 4, 4)
]
passed = True
for e, w in zip(targ_weights, uweights):
norm = np.linalg.norm(e-w[-1])
if norm > .000001:
passed = False
print((norm, e, w[-1]), file=sys.stderr)
self.assertIs(passed, True)
finite_difference
def test_finite_difference(self):
sin_grid = np.linspace(0, 2 * np.pi, 200)
sin_vals = np.sin(sin_grid)
deriv = finite_difference(sin_grid, sin_vals, 1) # 3rd deriv
sin_grid = np.arange(0, 1, .001)
sin_vals = np.sin(sin_grid)
sin_d3_vals = -np.cos(sin_grid)
if self.save_data:
f = h5py.File(TestManager.test_data('fd_data_1D.hdf5'),'w')
f.create_dataset("vals", data=sin_vals)
f.create_dataset("ref", data=sin_d3_vals)
print_error = False
plot_error = False
ref_vals = np.cos(sin_grid)
for ord in range(2, 10):
n = 1
sten = ord
# core_dvals = sin_d_vals[math.floor(sten/2):-math.floor(sten/2)]
vals = finite_difference(sin_grid, sin_vals, n, stencil=sten, end_point_accuracy=0,
mode="sparse"
)
errs = self.get_error(ref_vals, vals)
if plot_error:
self.plot_err(sin_grid, ref_vals, vals, errs)
if print_error:
self.print_error(n, ord, errs)
self.assertLess(errs[1], .1/ord)
ref_vals = -np.sin(sin_grid)
for ord in range(3, 10):
n=2
sten = ord
vals = finite_difference(sin_grid, sin_vals, n, stencil=sten, end_point_accuracy=0,
mode="dense"
)
errs = self.get_error(ref_vals, vals)
if plot_error:
self.plot_err(sin_grid, ref_vals, vals, errs)
if print_error:
self.print_error(n, ord, errs)
self.assertLess(errs[1], .05 / ord)
ref_vals = sin_d3_vals
for ord in range(4, 10):
n = 3
sten = ord
vals = finite_difference(sin_grid, sin_vals, n, ord, stencil=sten, end_point_accuracy=2,
mode="sparse"
)
errs = self.get_error(ref_vals, vals)
if plot_error:
self.plot_err(sin_grid, ref_vals, vals, errs)
if print_error:
self.print_error(n, ord, errs)
self.assertLess(errs[1], .05 / ord)
FD2D
def test_FD2D(self):
print_error = False
plot_error = False
x_grid = np.linspace(0, np.pi, 200, dtype=np.longdouble)
y_grid = np.linspace(0, np.pi, 100, dtype=np.longdouble)
sin_x_vals = np.sin(x_grid); sin_y_vals = np.sin(y_grid)
cos_x_vals = np.cos(x_grid); cos_y_vals = np.cos(y_grid)
vals_2D = np.outer(sin_x_vals, sin_y_vals)
grid_2D = np.array(np.meshgrid(x_grid, y_grid)).T
# x_grid_new = np.linspace(0, 2 * np.pi, 200)
# y_grid_new = np.linspace(0, 2 * np.pi, 100)
#
# sin_x_vals = np.sin(x_grid_new); sin_y_vals = np.sin(y_grid_new)
# vals_2D = np.outer(sin_x_vals, sin_y_vals)
# grid_2D = np.array(np.meshgrid(x_grid, y_grid)).T
#
# deriv = finite_difference(grid_2D, vals_2D, (1, 3))
#
#
# deriv = finite_difference(grid_2D, vals_2D, (1, 1))
# deriv = finite_difference(grid_2D, vals_2D, (1, 2))
# deriv = finite_difference(grid_2D, vals_2D, (1, 3))
# try 1st and 1st derivs
test_11 = True
if test_11:
ref_vals = np.outer(cos_x_vals, cos_y_vals)
for ord in range(3, 7, 2):
n = (1, 1)
vals = finite_difference(grid_2D, vals_2D, n, stencil=ord, end_point_accuracy=1)
errs = self.get_error(ref_vals, vals)
if plot_error:
self.plot_err(grid_2D, ref_vals, vals, errs)
if print_error:
self.print_error(n, ord, errs)
self.assertLess(errs[1], .1 / ord)
# 1,2 mixed deriv
test_12 = True
if test_12:
ref_vals = -np.outer(cos_x_vals, sin_y_vals)
for ord in range(3, 7, 2):
n = (1, 2)
vals = finite_difference(grid_2D, vals_2D, n, stencil=ord, end_point_accuracy=1)
errs = self.get_error(ref_vals, vals)
if plot_error:
self.plot_err(grid_2D, ref_vals, vals, errs)
if print_error:
self.print_error(n, ord, errs)
self.assertLess(errs[1], .05 / ord)
# 2,2 mixed deriv
test_23 = True
if test_23:
only_core = True
ref_vals = np.outer(sin_x_vals, cos_y_vals)
for ord in range(5, 10):
n = (2, 3)
vals = finite_difference(grid_2D, vals_2D, n,
stencil=ord,
end_point_accuracy=2,
only_core=True
)
floop = np.math.floor(ord/2)
if only_core:
refs = ref_vals[floop:-floop, floop:-floop]
grfs = grid_2D[floop:-floop, floop:-floop]
else:
refs = ref_vals
grfs = grid_2D
errs = self.get_error(refs, vals)
if plot_error:
self.plot_err(grfs, refs, vals, errs)
if print_error:
self.print_error(n, ord, errs)
self.assertLess(errs[1], .05)
# 1,4 mixed deriv
test_14 = True
if test_14:
ref_vals = np.outer(cos_x_vals, sin_y_vals)
only_core = True
for ord in range(6, 8):
n = (1, 4)
# vals = finite_difference(grid_2D, vals_2D, n,
# stencil=ord, end_point_accuracy=2,
# only_core = only_core
# )
vals = finite_difference(grid_2D, vals_2D, n,
stencil=ord, end_point_accuracy=2,
only_core = only_core
)
floop = np.math.floor(ord / 2)
if only_core:
refs = ref_vals[floop:-floop, floop:-floop]
grfs = grid_2D[floop:-floop, floop:-floop]
else:
refs = ref_vals
grfs = grid_2D
errs = self.get_error(refs, vals)
if plot_error:
self.plot_err(grfs, refs, vals, errs)
if print_error:
self.print_error(n, ord, errs)
self.assertLess(errs[1], .5)
FD2D_multi
def test_FD2D_multi(self):
print_error = False
plot_error = False
x_grid = np.arange(0, 1, .01, dtype=np.longdouble)
y_grid = np.arange(0, 1, .001, dtype=np.longdouble)
sin_x_vals = np.sin(x_grid); sin_y_vals = np.sin(y_grid)
cos_x_vals = np.cos(x_grid); cos_y_vals = np.cos(y_grid)
vals_2D = np.outer(sin_x_vals, sin_y_vals)
grid_2D = np.array(np.meshgrid(x_grid, y_grid)).T
# try 1st and 1st derivs
test_11 = True
if test_11:
ref_vals = np.outer(cos_x_vals, cos_y_vals)
for ord in range(3, 7, 2):
n = (1, 1)
nreps = 15
vals = finite_difference(
np.broadcast_to(grid_2D, (nreps,) + grid_2D.shape),
np.broadcast_to(vals_2D, (nreps,) + vals_2D.shape),
n,
stencil=ord,
end_point_accuracy=1,
contract = True
)
errs = self.get_error(ref_vals, vals)
if plot_error:
self.plot_err(grid_2D, ref_vals, vals, errs)
if print_error:
self.print_error(n, ord, errs)
self.assertLess(errs[1], .1 / ord)
FDDeriv
def test_FDDeriv(self):
# ggrid = GraphicsGrid(nrows=1, ncols=2)
fdf = FiniteDifferenceDerivative(np.sin, stencil = 5)
grid = np.linspace(0, 2*np.pi, 5)
diff_fun = fdf(grid, mesh_spacing = .1)
d = np.array([-.2, -.1, 0, .1, .2])
raw = np.array([
finite_difference(
c + d,
np.sin(c + d),
(1,),
stencil=5, only_center = True, end_point_accuracy=0
) for c in grid
])
self.assertLess(np.linalg.norm(diff_fun[0] - raw), .00001)
self.assertIsInstance(diff_fun[0, 0, 0][0], (float,))
FDDeriv2
def test_FDDeriv2(self):
def test(gps):
x = gps[..., 0]
y = gps[..., 1]
return x * np.sin(y)
fdf = FiniteDifferenceDerivative(test, function_shape=((2, ), 1), stencil = 5)
npts = 25
grid = np.linspace(0, 2*np.pi, npts)
grid2D = np.meshgrid(grid, grid)
gps = np.array(grid2D).T
diff_fun = fdf(gps)
diff_fun[0, 0]
self.assertEquals(diff_fun[0, 0].shape, (npts, npts))
self.assertEquals(diff_fun[0, :].shape, (1, 2, npts, npts)) # should I implement this with a squeeze...?
self.assertEquals(np.linalg.norm((diff_fun[:, :][1, 0] - diff_fun[1, 0]).flatten()), 0.)
FiniteDifferenceParallelism
def test_FiniteDifferenceParallelism(self):
re_1 = 0.9575
re_2 = 0.9575
b_e = np.deg2rad(104.5)
internals = [
[0, -1, -1, -1],
[1, 0, -1, -1],
[2, 0, 1, -1]
]
# internals = None
coords = np.array([
[0.000000, 0.000000, 0.000000],
[re_1, 0.000000, 0.000000],
np.dot(
nput.rotation_matrix([0, 0, 1], b_e),
[re_2, 0.000000, 0.000000]
)
])
erg2h = UnitsData.convert("Ergs", "Hartrees")
cm2borh = UnitsData.convert("InverseAngstroms", "InverseBohrRadius")
De_1 = 8.84e-12 * erg2h
a_1 = 2.175 * cm2borh
De_2 = 8.84e-12 * erg2h
a_2 = 2.175 * cm2borh
hz2h = UnitsData.convert("Hertz", "Hartrees")
kb = 7.2916e14 / (2 * np.pi) * hz2h
morse = self.harmonically_coupled_morse(
De_1, a_1, re_1,
De_2, a_2, re_2,
kb, b_e
)
deriv_gen = FiniteDifferenceDerivative(morse,
function_shape=((None, None), 0),
mesh_spacing=1e-3,
stencil=9,
logger=Logger(),
parallelizer=MultiprocessingParallelizer()#, verbose=True)
).derivatives(coords)
with BlockProfiler("With parallelizer"):
pot_derivs = deriv_gen.derivative_tensor([1, 2, 3, 4, 5, 6, 7])
deriv_gen = FiniteDifferenceDerivative(morse,
function_shape=((None, None), 0),
mesh_spacing=1e-3,
logger=Logger(),
stencil=9,
).derivatives(coords)
with BlockProfiler("Without parallelizer"):
pot_derivs = deriv_gen.derivative_tensor([1, 2, 3, 4, 5, 6])
TensorTerms
def test_TensorTerms(self):
n_Q = 10
n_X = 20
x_derivs = [
np.random.rand(n_Q, n_X),
np.random.rand(n_Q, n_Q, n_X),
np.random.rand(n_Q, n_Q, n_Q, n_X),
np.random.rand(n_Q, n_Q, n_Q, n_Q, n_X)
]
v_derivs = [
np.random.rand(n_X),
np.random.rand(n_X, n_X),
np.random.rand(n_X, n_X, n_X),
np.random.rand(n_X, n_X, n_X, n_X)
]
terms = TensorExpansionTerms(x_derivs, v_derivs)
q1 = terms.QX(1)
self.assertEquals(str(q1), 'Q[1]')
q11 = terms.QX(1) + terms.QX(1)
self.assertEquals(str(q11), 'Q[1]+Q[1]')
qshift = q11.shift(2, 1)
self.assertEquals(str(qshift), '[Q[1]+Q[1]|2->1]')
vQ = terms.QX(1).dot(terms.XV(1), 2, 1)
self.assertEquals(str(vQ), '<Q[1]:2,1:V[1]>')
self.assertIsInstance(vQ, TensorExpansionTerms.ContractionTerm)
# self.assertIsInstance(vQ.simplify(), TensorExpansionTerms.BasicContractionTerm)
vdQQ = vQ.dQ()
self.assertIn("V[2]", str(vdQQ))
self.assertIn("<Q[2]:3,1:V[1]>", str(vdQQ))
vdQQQQ = vdQQ.dQ().dQ().reduce_terms()
self.assertIsInstance(vdQQQQ, TensorExpansionTerms.SumTerm)
self.assertEquals(len(vdQQQQ.terms), 15)
# arr = vdQQQQ.array
xv_derivs = [
[None, np.random.rand(n_Q, n_X, n_X)],
[None, np.random.rand(n_Q, n_Q, n_X, n_X)]
]
mixed_terms = TensorExpansionTerms(x_derivs, v_derivs, qxv_terms=xv_derivs)
vdQ = mixed_terms.QX(1).dot(mixed_terms.XV(1), 2, 1)
mixed_vdQXX = vdQ.dQ().dQ().reduce_terms()
vdQQQQ_subbed = mixed_vdQXX.dQ().reduce_terms()
self.assertEquals(len(vdQQQQ_subbed.terms), 14)
self.assertEquals(vdQQQQ_subbed.array.shape, (n_Q, n_Q, n_Q, n_Q))
TensorConversion
def test_TensorConversion(self):
n_Q = 10
n_X = 20
np.random.seed(1)
x_derivs = [
np.random.rand(n_Q, n_X),
np.random.rand(n_Q, n_Q, n_X),
np.random.rand(n_Q, n_Q, n_Q, n_X),
np.random.rand(n_Q, n_Q, n_Q, n_Q, n_X)
]
v_derivs = [
np.random.rand(n_X),
np.random.rand(n_X, n_X),
np.random.rand(n_X, n_X, n_X),
np.random.rand(n_X, n_X, n_X, n_X)
]
xv_derivs = [
[None, np.random.rand(n_Q, n_X, n_X)],
[None, np.random.rand(n_Q, n_Q, n_X, n_X)]
]
t = TensorDerivativeConverter(x_derivs, v_derivs)
new = t.convert()
self.assertEquals(len(new), 4)
self.assertEquals(new[1].shape, (n_Q, n_Q))
t2 = TensorDerivativeConverter(x_derivs, v_derivs, mixed_terms=xv_derivs)
new = t2.convert()
self.assertEquals(len(new), 4)
self.assertEquals(new[1].shape, (n_Q, n_Q))
trace_derivs = TensorExpansionTerms([0, 0, 0, 0, 0], [0, 0, 0, 0, 0])
t1 = trace_derivs.QX(1).tr().dQ()
self.assertEquals(str(t1), 'Tr[Q[2],2+3]')
t1 = trace_derivs.QX(1).det().dQ().dQ()
t1 = trace_derivs.QX(1).inverse().dQ()
PseudopotentialTerms
def test_PseudopotentialTerms(self):
n_Q = 10
np.random.seed(1)
n_X = 3
i_derivs = [
np.random.rand(n_X, n_X),
np.random.rand(n_Q, n_X, n_X),
np.random.rand(n_Q, n_Q, n_X, n_X),
np.random.rand(n_Q, n_Q, n_Q, n_X, n_X)
]
# for x in i_derivs: # symmetrize
# inds = np.indices(x.shape).T.reshape(-1, x.ndim)
# for i in inds:
# n_X_part = i[-2:]
# n_Q_part = i[:-2]
# parent = np.concatenate([np.sort(n_Q_part), np.sort(n_X_part)])
# x[i] = x[parent]
i_terms = TensorExpansionTerms(i_derivs[1:], None, base_qx=i_derivs[0], q_name='I')
detI = i_terms.QX(0).det()
g_derivs = [
np.random.rand(n_Q, n_Q),
np.random.rand(n_Q, n_Q, n_Q),
np.random.rand(n_Q, n_Q, n_Q, n_Q),
np.random.rand(n_Q, n_Q, n_Q, n_Q, n_Q)
]
# for x in g_derivs: # symmetrize
# inds = np.indices(x.shape).T.reshape(-1, x.ndim)
# for i in inds:
# n_X_part = i[-2:]
# n_Q_part = i[:-2]
# parent = np.concatenate([np.sort(n_Q_part), np.sort(n_X_part)])
# x[i] = x[parent]
g_terms = TensorExpansionTerms(g_derivs[1:], None, base_qx=g_derivs[0], q_name='G')
detG = g_terms.QX(0).det()
self.assertIsInstance(detG.array, float)
self.assertEquals(detG.array, np.linalg.det(g_derivs[0]))
new_tr = g_terms.QX(0).tr()
self.assertEquals(new_tr.array.shape, ())
self.assertTrue(np.allclose(new_tr.array, np.trace(g_derivs[0])))
new_inv = ~g_terms.QX(0)
self.assertTrue(np.allclose(new_inv.array, np.linalg.inv(g_derivs[0])))
newdQ = detG.dQ()
self.assertEquals(newdQ.array.shape, (n_Q,))
i_derivs = [
np.random.rand(n_X, n_X),
np.random.rand(n_Q, n_X, n_X),
np.random.rand(n_Q, n_Q, n_X, n_X),
np.random.rand(n_Q, n_Q, n_Q, n_X, n_X)
]
# for x in i_derivs: # symmetrize
# inds = np.indices(x.shape).T.reshape(-1, x.ndim)
# for i in inds:
# n_X_part = i[-2:]
# n_Q_part = i[:-2]
# parent = np.concatenate([np.sort(n_Q_part), np.sort(n_X_part)])
# x[i] = x[parent]
i_terms = TensorExpansionTerms(i_derivs[1:], None, base_qx=i_derivs[0], q_name='I')
detI = i_terms.QX(0).det()
gam = detG / detI
self.assertEquals(gam.array.shape, ())
self.assertTrue(np.allclose(gam.array, detG.array / detI.array))
self.assertTrue(np.allclose(gam.array, np.linalg.det(g_derivs[0]) / np.linalg.det(i_derivs[0])))
five_gam = 5 * detG / detI
self.assertAlmostEquals(five_gam.array, 5 * detG.array / detI.array, 8)
inv_gam = 1/gam
self.assertEquals(inv_gam.array.shape, ())
self.assertTrue(np.allclose(inv_gam.array, detI.array / detG.array))
self.assertEquals(inv_gam.array, 1/gam.array)
gamdQ = gam.dQ().simplify(check_arrays=True)
self.assertEquals(gamdQ.array.shape, (n_Q,))
# self.assertEquals(gamdQQ.array.shape, (n_Q, n_Q))
# wat = g_terms.QX(0).dot(gamdQQ, [1, 2], [1, 2])
# self.assertEquals(wat.array.shape, ())
wat_2 = g_terms.QX(1).dot(gamdQ, 3, 1).tr()
self.assertEquals(wat_2.array.shape, ())
wat_21 = g_terms.QX(2).tr(axis1=1, axis2=4)
self.assertEquals(wat_21.array.shape, (n_Q, n_Q))
self.assertTrue(np.allclose(wat_21.array, np.trace(g_derivs[2], axis1=0, axis2=3)))
doot = gamdQ.dot(g_terms.QX(0), 1, 1)
self.assertEquals(doot.array.shape, (n_Q,))
wat_3 = -3 / 4 * gamdQ.dot(doot, 1, 1)
self.assertEquals(wat_3.array.shape, ())
wat_4 = -3 / 4 * inv_gam * gamdQ.dot(doot, 1, 1)
self.assertEquals(wat_4.array.shape, ())
wat_5 = inv_gam * wat_3
self.assertTrue(np.allclose(wat_4.array, wat_5.array))
I0, I0Q, I0QQ = i_derivs[:3]
G, GQ, GQQ = g_derivs[:3]
detI = np.linalg.det(I0); invI = np.linalg.inv(I0); adjI = invI * detI
detG = np.linalg.det(G); invG = np.linalg.inv(G); adjG = invG * detG
invIdQ_new = i_terms.QX(0).inverse().dQ()
invIdQ = - np.tensordot(np.tensordot(invI, I0Q, axes=[-1, 1]), invI, axes=[-1, 0]).transpose(1, 0, 2)
self.assertEquals(invIdQ_new.array.shape, invIdQ.shape)
self.assertTrue(np.allclose(invIdQ_new.array, invIdQ))
invGdQ = - np.tensordot(np.tensordot(invG, GQ, axes=[-1, 1]), invG, axes=[-1, 0]).transpose(1, 0, 2)
# not quite enough terms to want to be clever here...
nQ = GQ.shape[0]
## First derivatives of the determinant
detIdQ = np.array([
np.trace(np.dot(adjI, I0Q[i]))
for i in range(nQ)
])
detIdQ2 = np.trace(
np.tensordot(adjI, I0Q, axes=[1, 1]),
axis1=0,
axis2=2
)
detI_new = i_terms.QX(0).det()
detIdQ_new = detI_new.dQ()
self.assertTrue(np.allclose(detIdQ2, detIdQ))
self.assertEquals(detIdQ_new.array.shape, detIdQ.shape)
self.assertTrue(np.allclose(detIdQ_new.array, detIdQ))
detGdQ = np.array([
np.trace(np.dot(adjG, GQ[i]))
for i in range(nQ)
])
detG_new = g_terms.QX(0).det()
detGdQ_new = detG_new.dQ()
self.assertEquals(detGdQ_new.array.shape, detGdQ.shape)
self.assertTrue(np.allclose(detGdQ_new.array, detGdQ))
# two_dot = g_terms.QX(0).dot(gamdQQ, [1, 2], [1, 2])
# self.assertTrue(np.allclose(two_dot.array, np.tensordot(g_terms.QX(0).array, gamdQQ.array, axes=[[0, 1], [0, 1]])))
gamdQ = (detI_new.dQ()/detI_new + -1 * detG_new.dQ()/detG_new).simplify(check_arrays=True)
detI_new.name = "|I|"
gamdQ_I_new = (detI_new.dQ()/detI_new).simplify(check_arrays=True)
# raise Exception(gamdQ_I_new)
## Derivatives of ln Gamma
gamdQ_I = 1 / detI * detIdQ
gamdQ_G = 1 / detG * detGdQ
gamdQ_og = gamdQ_I - gamdQ_G
self.assertTrue(np.allclose(gamdQ.array, gamdQ_og))
adjIdQ = detI * invIdQ + detIdQ[:, np.newaxis, np.newaxis] * invI[np.newaxis, :, :]
np.array([
np.trace(np.dot(adjI, I0Q[i]))
for i in range(nQ)
])
detIdQQ = np.array([
[
np.tensordot(I0Q[i], adjIdQ[j].T, axes=2)
+ np.tensordot(adjI, I0QQ[i, j].T, axes=2)
for i in range(nQ)
]
for j in range(nQ)
])
detIdQQ_real = np.array([
[
np.trace(np.dot(I0Q[i], adjIdQ[j]))
+ np.trace(np.dot(adjI, I0QQ[i, j]))
for i in range(nQ)
]
for j in range(nQ)
])
detIdQQ2_terms = [
np.tensordot(I0Q, adjIdQ, axes=[[1, 2], [2, 1]]).T,
np.tensordot(adjI, I0QQ, axes=[[1, 0], [2, 3]]).T
]
detIdQQ2 = np.sum(detIdQQ2_terms, axis=0)
detIdQQ_new = detIdQ_new.dQ().simplify(check_arrays=True)
adjI_new = i_terms.QX(0).det() * i_terms.QX(0).inverse()
self.assertTrue(np.allclose(adjI_new.array, adjI))
adjIdQ_new = adjI_new.dQ().simplify()
self.assertTrue(np.allclose(adjIdQ_new.array, adjIdQ))
self.assertTrue(np.allclose(detIdQQ2, detIdQQ))
self.assertEquals(detIdQQ_new.array.shape, detIdQQ.shape)
self.assertTrue(np.allclose(detIdQQ_new.array.T, detIdQQ))
gamdQQ_I = -1 / detI ** 2 * np.outer(detIdQ, detIdQ) + 1 / detI * detIdQQ
gamdQQ_I_new = gamdQ_I_new.dQ().simplify(check_arrays=True)
# raise Exception(gamdQ_I_new, gamdQQ_I_new)
# raise Exception(gamdQQ_I_new.array.T, gamdQQ_I[0])
self.assertTrue(np.allclose(gamdQQ_I_new.array.T, gamdQQ_I))
ExplicitPoly
def test_ExplicitPoly(self):
a = FunctionExpansion.from_indices(
{
(): 1,
(0,): 2
},
ndim=2
)
self.assertTrue(np.allclose(
a([
[[1, 2], [.5, 2]],
[[1, 3], [.5, 3]],
[[1, 4], [.5, 4]]
]),
np.array([
[3, 2],
[3, 2],
[3, 2],
])
))
TensorDerivatives
def test_TensorDerivatives(self):
# a = TensorExpression.TermVector([TensorExpression.CoordinateTerm(0, 2), TensorExpression.CoordinateTerm(1, 2)])
def plus_1(x):
return x+1
crds = TensorExpression.ScalarFunctionTerm(
TensorExpression.CoordinateVector(2, name="coord_vec"),
f={'function':plus_1, 'derivatives':lambda inds: (lambda *_:1) if isinstance(inds, int) or len(inds) == 1 else (lambda *_:0)},
)
expr = TensorExpression.OuterPowerTerm(crds, 2)
e2 = TensorExpression.OuterPowerTerm(
TensorExpression.ScalarFunctionTerm(
TensorExpression.CoordinateVector([1, 2], name="coord_vec"),
f={'function': plus_1,
'derivatives': lambda inds: (lambda *_: 1) if isinstance(inds, int) or len(inds) == 1 else (
lambda *_: 0)},
), 2)
self.assertTrue(np.allclose(
TensorExpression(expr.dQ(), coord_vec=np.array([1, 2])).eval(),
e2.dQ().array
))
nv = TensorExpression.VectorNormTerm(TensorExpression.CoordinateVector(2, name="coord_vec"))
self.assertEquals(TensorExpression(nv, coord_vec=np.array([1, 2])).eval(), np.linalg.norm([1, 2]))
self.assertTrue(np.allclose(
TensorExpression(nv.dQ().dQ(), coord_vec=np.array([1, 2])).eval(),
[
[0.357771, -0.178885],
[-0.178885, 0.0894427]
]
))
np.random.seed(0)
crd = np.random.rand(5)
wat = TensorExpression.VectorNormTerm(
TensorExpression.CoordinateVector(5, name="coord_vec")
)
crd_vals = TensorExpression.ArrayStack((3,), np.array([crd, crd, crd]))
res = TensorExpression(wat, coord_vec=crd_vals).eval()
self.assertEquals(res.shape, (3,))
wat_d = wat.dQ()
dq_res = TensorExpression(wat_d, coord_vec=crd_vals).eval()
self.assertEquals(dq_res.shape, (3, 5))
slow_mode = np.array([
TensorExpression(wat_d, coord_vec=x).eval()
for x in crd_vals.array
])
self.assertTrue(np.allclose(dq_res, slow_mode))
wat_dd = wat.dQ().dQ()
dqq_res = TensorExpression(wat_dd, coord_vec=crd_vals).eval()
self.assertEquals(dqq_res.shape, (3, 5, 5))
slow_mode = np.array([
TensorExpression(wat_dd, coord_vec=x).eval()
for x in crd_vals.array
])
self.assertTrue(np.allclose(dqq_res, slow_mode))
pts = np.array([[-0.32498609, -0.29857954],
[-0.29112237, -0.48247142],
[-0.19237367, -0.28342343],
[-0.40490118, -0.2155546],
[-0.31110323, -0.10724408]])
pts = TensorExpression.ArrayStack((5,), pts)
term = TensorExpression.OuterPowerTerm(
TensorExpression.CoordinateVector(2, name="pts"),
2
)
# self.assertEquals(TensorExpression(term, pts=pts).eval().shape, (5, 2, 2))
self.assertEquals(TensorExpression(term.dQ(), pts=pts).eval().shape, (5, 2, 2, 2))
pts = np.array([[-0.32498609, -0.29857954],
[-0.29112237, -0.48247142],
[-0.19237367, -0.28342343],
[-0.40490118, -0.2155546],
[-0.31110323, -0.10724408]])
pts = TensorExpression.ArrayStack((5,), pts)
term = TensorExpression.OuterPowerTerm(
TensorExpression.CoordinateVector(2, name="pts"),
3
)
# self.assertEquals(TensorExpression(term, pts=pts).eval().shape, (5, 2, 2))
self.assertEquals(TensorExpression(term, pts=pts).eval().shape, (5, 2, 2, 2))
ExpandFunction
def test_ExpandFunction(self):
dtype = np.float32
def sin_xy(pt):
ax = -1 if pt.ndim > 1 else 0
return np.prod(np.sin(pt), axis=ax)
point = np.array([.5, .5], dtype=dtype)
exp = FunctionExpansion.expand_function(sin_xy, point, function_shape=((2,), 0), order=4, stencil=6)
hmm = np.vstack(
[np.linspace(-.5, .5, 100, dtype=dtype), np.zeros((100,), dtype=dtype)]
).T + point[np.newaxis]
# print(hmm)
ref = sin_xy(hmm)
test = exp(hmm)
plot_error = False
if plot_error:
exp2 = FunctionExpansion.expand_function(sin_xy, point, function_shape=((2,), 0), order=1, stencil=5)
g = hmm[:, 0]
gg = GraphicsGrid(nrows=1, ncols=2, tighten=True)
gg[0, 0] = Plot(g, exp(hmm), figure=Plot(g, sin_xy(hmm), figure=gg[0, 0]))
gg[0, 1] = Plot(g, exp2(hmm), figure=Plot(g, sin_xy(hmm), figure=gg[0, 1]))
gg.show()
plot2Derror = False
if plot2Derror:
# -0.008557147793556821113 0.007548466137326598213 0.0019501675856203966399
mesh = np.meshgrid(
np.linspace(.4, .6, 100, dtype=dtype),
np.linspace(.4, .6, 100, dtype=dtype)
)
grid = np.array(mesh).T
gg = GraphicsGrid(nrows=1, ncols=2)
ref2 = sin_xy(grid)
test2 = exp(grid)
err2 = ref2 - test2
# print(np.min(err2), np.max(err2), np.average(np.abs(err2)))
ContourPlot(*mesh, ref2 - test2, figure=gg[0, 1], plot_style=dict(vmin=-1e-4, vmax=1e-4))
ContourPlot(*mesh, test2, figure=gg[0, 0])
gg.show()
self.assertEquals(exp(point), exp.ref)
self.assertLess(np.linalg.norm(test - ref), .01)
ExpansionDerivs
def test_ExpansionDerivs(self):
dtype = np.float32
def sin_xy(pt):
ax = -1 if pt.ndim > 1 else 0
return np.prod(np.sin(pt), axis=ax)
point = np.array([.5], dtype=dtype)
disp = .01
exp1 = FunctionExpansion.expand_function(sin_xy, point, function_shape=((1,), 0), order=4, stencil=6)
poly_coeffs = np.array([exp1.expansion_tensors[i].flatten() for i in range(4)]).flatten()
dpoly_coeffs = np.array([exp1.deriv().expansion_tensors[i].flatten() for i in range(3)]).flatten()
dpoly2_coeffs = np.array([exp1.deriv().deriv().expansion_tensors[i].flatten() for i in range(2)]).flatten()
self.assertTrue(dpoly_coeffs[0] == 2*poly_coeffs[1])
self.assertTrue(dpoly2_coeffs[0] == 2 * 3 * poly_coeffs[2])
self.assertTrue(dpoly2_coeffs[1] == 3 * 4 * poly_coeffs[3])
def sin_xy_d(pt):
return np.array([
np.cos(pt[..., 0]) * np.sin(pt[..., 1]),
np.sin(pt[..., 0]) * np.cos(pt[..., 1])
]).T
exp2D = FunctionExpansion.expand_function(sin_xy, np.array([.5, -.5]),
function_shape=((2,), 0),
order=7,
stencil=11)
self.assertLess(
np.linalg.norm(
exp2D.deriv()([
exp2D.center,
exp2D.center - disp,
exp2D.center + disp,
]) -
sin_xy_d(np.array([
exp2D.center,
exp2D.center - disp,
exp2D.center + disp,
]))
),
.005)
MultiExpansion
def test_MultiExpansion(self):
dtype = np.float32
def sin_xy(pt):
ax = -1 if pt.ndim > 1 else 0
return np.prod(np.sin(pt), axis=ax)
point = np.array([.5, .5], dtype=dtype)
disp = .01
exp1 = FunctionExpansion.expand_function(sin_xy, point, function_shape=((2,), 0), order=4, stencil=6)
exp2 = FunctionExpansion.expand_function(sin_xy, point-disp, function_shape=((2,), 0), order=4, stencil=6)
exp3 = FunctionExpansion.expand_function(sin_xy, point+disp, function_shape=((2,), 0), order=4, stencil=6)
exp4 = FunctionExpansion.expand_function(sin_xy, point+2*disp, function_shape=((2,), 0), order=4, stencil=6)
multi = FunctionExpansion.multipolynomial(exp1, exp2, exp3, exp4)
d1 = exp1.deriv()
exp1([exp1.center, exp2.center, exp3.center, exp4.center])
d1([exp1.center, exp2.center, exp3.center, exp4.center])
mult_test_1 = multi([exp1.center, exp2.center, exp3.center, exp4.center])
mult_test_2 = multi([exp1.center, exp2.center, exp3.center, exp4.center], outer=False)
self.assertTrue(np.allclose(mult_test_2, np.diag(mult_test_1)))
Polys
def test_Polys(self):
d = SparsePolynomial({
#1 + 2x + 3y^2 - xy
():1,
(0,):2,
(1, 1):3,
(0, 1):-1
}).as_dense()
self.assertTrue(
np.allclose(d.coeffs, np.array([[1, 0, 3], [2, -1, 0]]))
)
self.assertTrue(
np.allclose(d.shift([1, 1]).coeffs, np.array([[5, 5, 3], [1, -1, 0]]))
)
d2 = SparsePolynomial({
#2 + y - 3x^2
():2,
(1,):1,
(0, 0):-3
}).as_dense()
d3 = SparsePolynomial({
# x - y
(0,): 1,
(1,): -1
}).as_dense()
multi = DensePolynomial(
[
np.pad(d.coeffs, [[0, 1], [0, 0]]),
np.pad(d2.coeffs, [[0, 0], [0, 1]])
],
stack_dim=1
)
# print(multi.coeffs)
# print(multi.deriv(1).coeffs)
# print(d.coefficient_tensors)
# print(d2.coefficient_tensors)
for n,(c1, c2, cm) in enumerate(zip(
d.coefficient_tensors,
d2.coefficient_tensors,
multi.coefficient_tensors
)):
self.assertTrue(
np.allclose(np.array([c1, c2]), cm)
)
# print(multi.coefficient_tensors)
multi_2 = DensePolynomial.from_tensors(multi.coefficient_tensors)
self.assertTrue(np.allclose(multi.coeffs, multi_2.coeffs))
for n, (c1, c2, cm) in enumerate(zip(
d.shift([1, 1]).coefficient_tensors,
d2.shift([2, 2]).coefficient_tensors,
multi.shift([[1, 1], [2, 2]]).coefficient_tensors
)):
self.assertTrue(
np.allclose(np.array([c1, c2]), cm)
)
for n, (c1, c2, cm) in enumerate(zip(
d.deriv(1).coefficient_tensors,
d2.deriv(1).coefficient_tensors,
multi.deriv(1).coefficient_tensors
)):
self.assertTrue(
np.allclose(np.array([c1, c2]), cm)
)
dg = d.grad()
for n, (c1, c2, cm) in enumerate(zip(
d.deriv(0).coefficient_tensors,
d.deriv(1).coefficient_tensors,
dg.coefficient_tensors
)):
self.assertTrue(
np.allclose(np.array([c1, c2]), cm)
)
pad_d3 = DensePolynomial(np.broadcast_to(d3.coeffs[np.newaxis], (2,) + d3.shape), stack_dim=1)
# raise Exception((multi*pad_d3).coeffs.shape, (d*d3).coeffs.shape)
for n, (c1, c2, cm) in enumerate(zip(
(d*d3).coefficient_tensors,
(d2*d3).coefficient_tensors,
(multi*pad_d3).coefficient_tensors
)):
self.assertTrue(
np.allclose(np.array([c1, c2]), cm)
)
SparseShifts
def test_SparseShifts(self):
d = SparsePolynomial({
# 1 + 2x + 3y^2 - xy
(): 1,
(0,): 2,
(1, 1): 3,
(0, 1): -1
},
ndim=2
)
print(d.terms)
shift_poly = d.shift([0, 1]).as_dense()
print(d.as_dense().coeffs)
print(shift_poly.coeffs)
print(d.as_dense().shift([0, 1]).coeffs)
SparseArrayPoly
def test_SparseArrayPoly(self):
from McUtils.Numputils import SparseArray
shape = (10, 3, 2, 1)
n_els = 3
np.random.seed(1)
inds = np.unique(np.array([np.random.choice(x, n_els) for x in shape]).T, axis=0)
vals = np.random.rand(len(inds))
inds = inds.T
# `from_data` for backend flexibility
array = SparseArray.from_data(
(
vals,
inds
),
shape=shape
)
poly = DensePolynomial(array, stack_dim=1)
doly = DensePolynomial(array.asarray(), stack_dim=1)
self.assertTrue(
np.allclose(
(poly + poly).coeffs.asarray(),
(doly + doly).coeffs
)
)
shape = (10, 4, 3, 7)
n_els = 4
np.random.seed(1)
inds = np.unique(np.array([np.random.choice(x, n_els) for x in shape]).T, axis=0)
vals = np.random.rand(len(inds))
inds = inds.T
# `from_data` for backend flexibility
ar2 = SparseArray.from_data(
(
vals,
inds
),
shape=shape
)
goly = DensePolynomial(ar2, stack_dim=1)
dggy = DensePolynomial(ar2.asarray(), stack_dim=1)
# print((poly + goly).coeffs.asarray())
# print((doly + dggy).coeffs)
self.assertTrue(
np.allclose(
(poly + goly).coeffs.asarray(),
(doly + dggy).coeffs
)
)
# goly.grad()
# print(poly.grad().shape)
# print(poly.grad().coeffs.asarray())
# print(doly.grad().coeffs)
self.assertTrue(
np.allclose(
poly.grad().coeffs.asarray(),
doly.grad().coeffs
)
)
self.assertTrue(
np.allclose(
(poly + goly).grad().coeffs.asarray(),
(doly + dggy).grad().coeffs
)
)
SparseConvolve
def test_SparseConvolve(self):
from McUtils.Numputils import SparseArray
a1 = self.constructSparseArray((3, 2), 3)
a2 = self.constructSparseArray((2, 5), 2)
p3 = DensePolynomial(a1.asarray()) * DensePolynomial(a2.asarray())
p4 = DensePolynomial(a1) * DensePolynomial(a2)
self.assertTrue(
np.allclose(p3.coeffs, p4.coeffs.asarray())
)
a1 = self.constructSparseArray((5, 3, 5), 4)
a2 = self.constructSparseArray((5, 8, 3), 7)
p3 = DensePolynomial(a1.asarray(), stack_dim=1) * DensePolynomial(a2.asarray(), stack_dim=1)
p4 = DensePolynomial(a1, stack_dim=1) * DensePolynomial(a2, stack_dim=1)
self.assertTrue(
np.allclose(p3.coeffs, p4.coeffs.asarray())
)
a1 = self.constructSparseArray((5, 3, 5, 2), 2)
a2 = self.constructSparseArray((5, 2, 3), 7)
p3 = DensePolynomial(a1.asarray(), stack_dim=1) + DensePolynomial(a2.asarray(), stack_dim=1)
p4 = DensePolynomial(a1, stack_dim=1) + DensePolynomial(a2, stack_dim=1)
self.assertTrue(
np.allclose(p3.coeffs, p4.coeffs.asarray())
)
p3 = DensePolynomial(a1.asarray(), stack_dim=1) * DensePolynomial(a2.asarray(), stack_dim=1)
p4 = DensePolynomial(a1, stack_dim=1) * DensePolynomial(a2, stack_dim=1)
self.assertTrue(
np.allclose(p3.coeffs, p4.coeffs.asarray())
)
LinSpaceMesh
def test_LinSpaceMesh(self):
mg = Mesh(np.linspace(-1, 1, 3))
self.assertIs(mg.mesh_type, MeshType.Structured)
self.assertEquals(mg.shape, (3,))
MeshGridMesh
def test_MeshGridMesh(self):
mg = np.array(np.meshgrid(np.array([-1, 0, 1]), np.array([-1, 0, 1]), indexing='ij'))
regmesh = Mesh(mg)
self.assertIs(regmesh.mesh_type, MeshType.Regular)
self.assertEquals(regmesh.shape, (3, 3, 2))
StructuredMesh
def test_StructuredMesh(self):
mg = np.array(np.meshgrid(np.array([-1, -.5, 0, 1]), np.array([-1, 0, 1]), indexing='ij'))
regmesh = Mesh(mg)
self.assertIs(regmesh.mesh_type, MeshType.Structured)
self.assertEquals(regmesh.shape, (4, 3, 2))
UnstructuredMesh
def test_UnstructuredMesh(self):
mg = np.random.rand(100, 10, 2)
regmesh = Mesh(mg)
self.assertIs(regmesh.mesh_type, MeshType.Unstructured)
RegularMesh
def test_RegularMesh(self):
regmesh = Mesh.RegularMesh([-1, 1, 3], [-1, 1, 3])
self.assertIs(regmesh.mesh_type, MeshType.Regular)
self.assertEquals(regmesh.shape, (3, 3, 2))
RegMeshSubgrids
def test_RegMeshSubgrids(self):
regmesh = Mesh.RegularMesh([-1, 1, 3], [-1, 1, 3])
m = [Mesh(g) for g in regmesh.subgrids]
self.assertTrue(all(g.mesh_type is MeshType.Regular for g in m))
SemiStructuredMesh
def test_SemiStructuredMesh(self):
semi_mesh = Mesh([[np.arange(i)] for i in range(10, 25)])
self.assertIs(semi_mesh.mesh_type, MeshType.SemiStructured)
MeshFromList
def test_MeshFromList(self):
try:
wat = np.asarray([[-1, 0, 1], [-1, 0, 1]])
bad = Mesh(wat)
except Mesh.MeshError:
pass
else:
# just to make it accessible through Mesh
self.assertIs(bad.mesh_type, MeshType.Indeterminate)
ParametricCurveInterpolation
def test_ParametricCurveInterpolation(self):
grid = np.linspace(0, 1, 8)
vals = np.array([
np.cos(grid),
np.sin(grid),
np.sin(grid) + np.cos(grid)
]).T
interp = Interpolator(grid, vals)
print(interp(.5))
RegularGridInterpolatorND
def test_RegularGridInterpolatorND(self):
cos_grid = np.linspace(0, 1, 8)
sin_grid1 = np.linspace(0, 1, 11)
sin_grid = np.linspace(0, 1, 12)
grids = [sin_grid, sin_grid1, cos_grid]
x, y, z = np.meshgrid(*grids, indexing='ij')
sin_vals = np.sin(x) + np.sin(y) + np.cos(z)
reg_interp = ProductGridInterpolator(grids, sin_vals, order=(3, 2, 4))
test_points = np.reshape(np.moveaxis(np.array([x, y, z]), 0, 3), (-1, 3))
vals = reg_interp(test_points)
self.assertTrue(np.allclose(vals, np.reshape(sin_vals, -1)))
test_points = np.random.uniform(0, 1, size=(100, 3))
test_vals = np.sin(test_points[:, 0]) + np.sin(test_points[:, 1]) + np.cos(test_points[:, 2])
interp_vals = reg_interp(test_points)
self.assertTrue(np.allclose(interp_vals, test_vals, atol=5e-5))
deriv = reg_interp.derivative((2, 0, 0))
test_deriv_vals = -np.sin(test_points[:, 0])# + np.sin(test_points[:, 1]) + np.cos(test_points[:, 2])
interp_vals = deriv(test_points)
self.assertTrue(np.allclose(interp_vals, test_deriv_vals, atol=5e-3))
cos_grid = np.linspace(0, 1, 8)
sin_grid1 = np.linspace(0, 1, 11)
sin_grid = np.linspace(0, 1, 12)
grids = [sin_grid, sin_grid1, cos_grid]
sin_vals = np.sin(x) * np.sin(y) * np.cos(z)
reg_interp = ProductGridInterpolator(grids, sin_vals)
deriv = reg_interp.derivative((1, 1, 0))
test_deriv_vals = np.cos(test_points[:, 0]) * np.cos(test_points[:, 1]) * np.cos(test_points[:, 2])
interp_vals = deriv(test_points)
self.assertTrue(np.allclose(interp_vals, test_deriv_vals, atol=5e-4))
Interpolator1D
def test_Interpolator1D(self):
def test_fn(grid):
return np.sin(grid)
sin_grid = np.linspace(0, 1, 12)
grid = sin_grid
sin_vals = test_fn(sin_grid)
interp = Interpolator(grid, sin_vals)
test_points = np.random.uniform(0, 1, size=(100,))
test_vals = test_fn(test_points)
interp_vals = interp(test_points)
self.assertTrue(np.allclose(interp_vals, test_vals, atol=5e-5))
InterpolatorExtrapolator2D
def test_InterpolatorExtrapolator2D(self):
def test_fn(grid):
return (
.25 * (np.sin(grid[..., 0])**2) * np.cos(grid[..., 1])
+ .5 * np.sin(grid[..., 0])*np.cos(grid[..., 1])
+ .25 * np.sin(grid[..., 0]) * (np.cos(grid[..., 1])**2)
)
cos_grid = np.linspace(0, np.pi, 8)
sin_grid = np.linspace(0, np.pi, 11)
grids = [sin_grid, cos_grid]
grid = np.moveaxis(np.array(np.meshgrid(*grids, indexing='ij')), 0, 2)
vals = test_fn(grid)
interp = Interpolator(grid, vals)
lin_ext_interp = Interpolator(grid, vals, extrapolator=Interpolator.DefaultExtrapolator, extrapolation_order=1)
g = GraphicsGrid(nrows=1, ncols=3,
figure_label='Extrapolation tests',
spacings=(70, 0)
)
g.padding_top = 40
g[0, 0].plot_label = "Analytic"
g[0, 1].plot_label = "Cubic Extrapolation"
g[0, 2].plot_label = "Linear Extrapolation"
big_grid = np.array(np.meshgrid(
np.linspace(-np.pi/2, 3*np.pi/2, 500),
np.linspace(-np.pi/2, 3*np.pi/2, 500)
))
eval_grid = np.moveaxis(big_grid, 0, 2)#.transpose((1, 0, 2))
inner_grid = np.array(np.meshgrid(
np.linspace(0, np.pi, 500),
np.linspace(0, np.pi, 500)
))
inner_eval_grid = np.moveaxis(inner_grid, 0, 2)
surf = test_fn(eval_grid)
ContourPlot(*big_grid, surf, figure=g[0, 0],
plot_style=dict(vmin=np.min(surf), vmax=np.max(surf))
)
ContourPlot(*big_grid, interp(eval_grid), figure=g[0, 1],
plot_style=dict(vmin=np.min(surf), vmax=np.max(surf)))
ContourPlot(*inner_grid, interp(inner_eval_grid), figure=g[0, 1],
plot_style=dict(vmin=np.min(surf), vmax=np.max(surf)))
ContourPlot(*big_grid, lin_ext_interp(eval_grid), figure=g[0, 2],
plot_style=dict(vmin=np.min(surf), vmax=np.max(surf)))
ContourPlot(*inner_grid, lin_ext_interp(inner_eval_grid), figure=g[0, 2],
plot_style=dict(vmin=np.min(surf), vmax=np.max(surf)))
ScatterPlot(*np.moveaxis(grid, 2, 0), figure=g[0, 0],
plot_style=dict(color='red')
)
ScatterPlot(*np.moveaxis(grid, 2, 0), figure=g[0, 1],
plot_style=dict(color='red')
)
ScatterPlot(*np.moveaxis(grid, 2, 0), figure=g[0, 2],
plot_style=dict(color='red')
)
g.show()
InterpolatorExtrapolator1D
def test_InterpolatorExtrapolator1D(self):
sin_grid = np.linspace(0, np.pi, 5)
sin_vals = np.sin(sin_grid)
interp = Interpolator(sin_grid, sin_vals,
extrapolator=Interpolator.DefaultExtrapolator, extrapolation_order=1)
default_interp = Interpolator(sin_grid, sin_vals)
test_points = np.random.uniform(-np.pi, 2*np.pi, size=(20,))
test_vals = np.sin(test_points)
interp_vals = interp(test_points)
RegularInterpolator3D
def test_RegularInterpolator3D(self):
def test_fn(grid):
return np.sin(grid[..., 0])*np.cos(grid[..., 1]) - np.sin(grid[..., 2])
cos_grid = np.linspace(0, 1, 8)
sin_grid1 = np.linspace(0, 1, 11)
sin_grid = np.linspace(0, 1, 12)
grids = [sin_grid, sin_grid1, cos_grid]
grid = np.moveaxis(np.array(np.meshgrid(*grids, indexing='ij')), 0, 3)
vals = test_fn(grid)
interp = Interpolator(grid, vals)
test_points = np.random.uniform(0, 1, size=(100000, 3))
test_vals = test_fn(test_points)
interp_vals = interp(test_points)
self.assertTrue(np.allclose(interp_vals, test_vals, atol=5e-5))
IrregularInterpolator3D
def test_IrregularInterpolator3D(self):
def test_fn(grid):
return np.sin(grid[..., 0])*np.cos(grid[..., 1]) - np.sin(grid[..., 2])
np.random.seed(3)
grid = np.random.uniform(0, 1, size=(10000, 3))
vals = test_fn(grid)
interp = Interpolator(grid, vals)
test_points = np.random.uniform(0, 1, size=(1000, 3))
test_vals = test_fn(test_points)
interp_vals = interp(test_points)
self.assertTrue(np.allclose(interp_vals, test_vals, atol=5e-3), msg='max diff: {}'.format(
np.max(np.abs(interp_vals - test_vals))
))
TensorExpressionEfficiency
def test_TensorExpressionEfficiency(self):
te = TensorExpression.OuterPowerTerm(TensorExpression.ConstantArray([1, 2, 3], name="bloop"), 3)
tet2 = TensorExpression.OuterTerm(
TensorExpression.OuterPowerTerm(TensorExpression.ConstantArray([1, 2, 3], name="bloop"), 2),
TensorExpression.ConstantArray([1, 2, 3], name="bloop").dQ()
)
c={te.dQ().terms[1]:1, tet2:2}
raise Exception(c)
TensorExpression.OuterPowerTerm(TensorExpression.ConstantArray([1, 2, 3], name="bloop"), 2)
RBFInterp1D
def test_RBFInterp1D(self):
# 1D
np.random.seed(0)
npts = 50
ndim = 1
pts = np.random.uniform(size=(npts, ndim))
vals = np.product(np.sin(pts), axis=1)
dvals_x = np.cos(pts[:, 0])
d2vals_x = -np.sin(pts[:, 0])
interp = RBFDInterpolator(
pts,
vals,
np.array([dvals_x]).T,
np.reshape(d2vals_x, (npts, ndim, ndim)),
kernel='thin_plate_spline'
)
pts = interp.grid
vals = interp(pts[:5], deriv_order=0, neighbors=5)
true = np.product(np.sin(pts[:5]), axis=1)
self.assertTrue(
np.allclose(
vals,
true
), msg="bad interpolation at interpolation points \n{} \nvs\n{}".format(
vals,
true
)
)
dervs = interp(pts[:2], deriv_order=1, neighbors=5)[1]
true = np.array([dvals_x]).T[:2]
self.assertTrue(
np.allclose(
dervs,
true
), msg="bad deriv interpolation at interpolation points \n{} \nvs\n {}".format(
dervs,
true
)
)
self.assertLess(
np.linalg.norm(
interp(pts[:2] + .05, deriv_order=0, neighbors=5) -
np.product(np.sin(pts[:2]), axis=1)
),
.2)
RBFInterpolator
def test_RBFInterpolator(self):
#
np.random.seed(1)
npts = 1000
ndim = 2
pts = np.random.uniform(low=-np.pi/2, high=np.pi/2, size=(npts, ndim))
og = pts
vals = og_vals = np.product(np.sin(pts), axis=1)
dvals_x = np.sin(pts[:, 1])*np.cos(pts[:, 0])
dvals_y = np.sin(pts[:, 0])*np.cos(pts[:, 1])
dvals = np.array([dvals_x, dvals_y]).T
og_d = dvals
dvals_xx = -np.sin(pts[:, 0])*np.sin(pts[:, 1])
dvals_xy = np.cos(pts[:, 1])*np.cos(pts[:, 0])
dvals_yy = -np.sin(pts[:, 1])*np.sin(pts[:, 0])
d2vals = np.moveaxis(np.array([[dvals_xx, dvals_xy], [dvals_xy, dvals_yy]]), -1, 0)
og_dd = d2vals
# import McUtils.Plots as plt
#
# plt.ScatterPlot3D(*pts.T, vals, plot_range=[[-np.pi/2, np.pi/2], [-np.pi/2, np.pi/2], [-1, 1]]).show()
interp = RBFDInterpolator(
pts,
vals,
dvals,
d2vals,
# extra_degree=2,
kernel='thin_plate_spline',
clustering_radius=.01,
multicenter_monomials=True,
# monomial_basis=True
)
pts = interp.grid
vals = np.product(np.sin(pts), axis=1)
dvals_x = np.sin(pts[:, 1]) * np.cos(pts[:, 0])
dvals_y = np.sin(pts[:, 0]) * np.cos(pts[:, 1])
dvals = np.array([dvals_x, dvals_y]).T
dvals_xx = -np.sin(pts[:, 0]) * np.sin(pts[:, 1])
dvals_xy = np.cos(pts[:, 1]) * np.cos(pts[:, 0])
dvals_yy = -np.sin(pts[:, 1]) * np.sin(pts[:, 0])
d2vals = np.moveaxis(np.array([[dvals_xx, dvals_xy], [dvals_xy, dvals_yy]]), -1, 0)
# raise Exception("???")
# with np.printoptions(linewidth=1e8, threshold=1e8):
# print("???", intl.matrix(pts[:1]))
test_vals = interp(pts[:2], neighbors=15)
real = np.product(np.sin(pts[:2]), axis=1)
self.assertTrue(np.allclose(
test_vals,
real,
atol=1e-3,
rtol=1e-2
), msg="bad interpolation at interpolation points \n {} \nvs\n {}".format(
test_vals, real
))
h = .001
test_pts = pts[:3] + np.array([[0, h]]*3)
extrap = interp(test_pts, neighbors=15, zero_tol=-1)
true = np.product(np.sin(test_pts), axis=1)
# if not np.allclose(
# extrap,
# true,
# atol=h
# ):
# plt.ScatterPlot3D(*test_pts.T, true,
# plot_range=[[-np.pi / 2, np.pi / 2], [-np.pi / 2, np.pi / 2], [-1, 1]]
# # figure=plt.ScatterPlot3D(*pts.T, vals)
# ).show()
self.assertTrue(
np.allclose(
extrap,
true,
atol=h
),
msg="bad extrapolation at pts: {} \nerr: {} in \n{} \nvs\n {}".format(
test_pts, extrap-true, extrap, true
)
)
test_pts = pts[:10]
dervs = interp(test_pts, neighbors=15, deriv_order=1, zero_tol=-1)[1]
reals = np.array(
[
dvals_x,
dvals_y
]).T[:10]
test_vals = vals[:10]
# if not np.allclose(
# dervs,
# reals,
# atol=.05
# ):
# plt.ScatterPlot3D(*test_pts.T, test_vals,
# plot_range=[[-np.pi / 2, np.pi / 2], [-np.pi / 2, np.pi / 2], [-1, 1]]
# ).show()
# plt.ScatterPlot3D(*pts[:3].T, test_vals).show()
self.assertTrue(np.allclose(
dervs,
reals,
atol=.1
), msg="bad deriv interpolation at interpolation points \nerror: {} in\n{} \nvs\n {}".format(dervs-reals, dervs, reals))
np.random.seed(0)
test_vals = np.unique(np.random.randint(0, len(pts)-1, size=200))
errors = [[], [], []]
print_errors = False
# prob_n = 8
# c = pts[test_vals[prob_n:prob_n+1]]+.1
#
# intl = interp.nearest_interpolation(c[0], neighbors=15, solver_data=True, interpolator=True)
# idat = intl.data
# import json, os
# with open(os.path.expanduser("~/Desktop/bad_pos_new.json"), 'w+') as dump:
# json.dump({
# 'center': c.tolist(),
# 'pts': pts.tolist(),
# 'vals': vals.tolist(),
# 'dvals': dvals.tolist(),
# 'd2vals': d2vals.tolist(),
# 'matrix': idat.solver_data[0].tolist(),
# 'weights': idat.weights[0].tolist(),
# 'centers': idat.scaling_data.reverse_renormalization(idat.centers).tolist(),
# 'test_mat': intl.matrix(pts[:1] + [0, .01], deriv_order=2).tolist()
# }, dump)
# raise Exception("...")
# print(dat.solver_data[0])
# raise Exception(dat.solver_data[0])
#
# raise Exception(
# interp(c, neighbors=15, reshape_derivatives=False),
# # interp.global_interpolator(c, reshape_derivatives=False),
# np.sin(c[:, 0]) * np.sin(c[:, 1])
# )
h = .1
for ifun in [
# interp.global_interpolator,
lambda x,**kw:interp(x, neighbors=25, **kw)
]:
for n,p in enumerate(pts[test_vals]):
c = p[np.newaxis]+h
test = ifun(c, reshape_derivatives=False, deriv_order=2)
# if np.abs(test[0])[0] > 1000:
# raise Exception(n)
real = [
np.sin(c[:, 0]) * np.sin(c[:, 1]),
np.array([
np.sin(c[:, 1]) * np.cos(c[:, 0]),
np.sin(c[:, 0]) * np.cos(c[:, 1])
]).T,
np.array([
-np.sin(c[:, 0]) * np.sin(c[:, 1]),
np.cos(c[:, 1]) * np.cos(c[:, 0]),
-np.sin(c[:, 1]) * np.sin(c[:, 0])
]).T
]
if print_errors:
print("-"*20+" ", n, ": ", c, " "+"-"*20)
for n, (t, r) in enumerate(zip(test, real)):
rel_error = 2*(t-r)/(np.abs(t)+np.abs(r))
if print_errors:
print(t, r, t-r)
print(">", rel_error)
errors[n].append(rel_error) # relative errors
maes = [np.average(np.abs(x), axis=None) for x in errors]
# print(maes)
tols = [h, 3*h, 10*h] # ruined by a single outlier...
for n,(t,e) in enumerate(zip(tols, maes)):
self.assertLess(e, t, msg="at order {} MRE {} > {}".format(n, e, t))
RBFInterpolatorResiliance
def test_RBFInterpolatorResiliance(self):
#
np.random.seed(1)
npts = 1000
ndim = 2
pts = np.random.uniform(low=-np.pi / 2, high=np.pi / 2, size=(npts, ndim))
og = pts
vals = og_vals = np.product(np.sin(pts), axis=1)
dvals_x = np.sin(pts[:, 1]) * np.cos(pts[:, 0])
dvals_y = np.sin(pts[:, 0]) * np.cos(pts[:, 1])
dvals = np.array([dvals_x, dvals_y]).T
og_d = dvals
dvals_xx = -np.sin(pts[:, 0]) * np.sin(pts[:, 1])
dvals_xy = np.cos(pts[:, 1]) * np.cos(pts[:, 0])
dvals_yy = -np.sin(pts[:, 1]) * np.sin(pts[:, 0])
d2vals = np.moveaxis(np.array([[dvals_xx, dvals_xy], [dvals_xy, dvals_yy]]), -1, 0)
og_dd = d2vals
# import McUtils.Plots as plt
#
# plt.ScatterPlot3D(*pts.T, vals, plot_range=[[-np.pi/2, np.pi/2], [-np.pi/2, np.pi/2], [-1, 1]]).show()
interp = RBFDInterpolator(
pts,
vals,
dvals,
d2vals,
# extra_degree=2,
kernel='thin_plate_spline',
clustering_radius=.01,
multicenter_monomials=True,
# monomial_basis=True
)
test_vals = interp(pts, neighbors=15, zero_tol=-1, resiliance_test_options={})
HigherElementaryDerivs
def test_HigherElementaryDerivs(self):
sym = Symbols('x', 'y')
fn = sym.morse(sym.x, de=2, a=1) + sym.morse(sym.y, de=2, a=1)
fexpr = 2*(1-sym.exp(-1*sym.x))**2 + 2*(1-sym.exp(-1*sym.y))**2
# raise Exception(sym.x + (3*sym.x**2)/12)
np.testing.assert_allclose(
fn.deriv(order=3)([[1, 2], [3, 4]]),
fexpr.deriv(order=3)([[1, 2], [3, 4]])
)
RBFTiming
def test_RBFTiming(self):
# for npts in [50, 100, 200, 300, 400, 500]:
# np.random.seed(1)
# npts = npts
# ndim = 2
# pts = np.random.uniform(size=(npts, ndim))
# vals = np.product(np.sin(pts), axis=1)
# dvals_x = np.sin(pts[:, 1])*np.cos(pts[:, 0])
# dvals_y = np.sin(pts[:, 0])*np.cos(pts[:, 1])
# dvals_xx = -np.sin(pts[:, 0])*np.sin(pts[:, 1])
# dvals_xy = np.cos(pts[:, 1])*np.cos(pts[:, 0])
# dvals_yy = -np.sin(pts[:, 1])*np.sin(pts[:, 0])
#
# interp = RBFDInterpolator(
# pts,
# vals,
# np.array([dvals_x, dvals_y]).T,
# np.moveaxis(np.array([[dvals_xx, dvals_xy], [dvals_xy, dvals_yy]]), -1, 0),
# extra_degree=2,
# kernel='gaussian',
# clustering_radius=.05
# )
#
# from Peeves.Timer import Timer
# with Timer(tag="{} pts:".format(len(interp.grid))):
# interp(interp.grid + .1, deriv_order=0, neighbors=5)
for npts in [300]:
np.random.seed(1)
npts = npts
ndim = 9
pts = np.random.uniform(size=(npts, ndim))
vals = np.product(np.sin(pts), axis=1)
dvals = []
for i in range(ndim):
vals = 1
for k in range(ndim):
if k != i:
vals = vals * np.sin(pts[:, k])
else:
vals = vals * np.cos(pts[:, k])
dvals.append(vals)
dvals = np.moveaxis(np.array(dvals), -1, 0)
d2vals = []
for i in range(ndim):
for j in range(ndim):
vals = 1
for k in range(ndim):
if k != i and k != j:
vals = vals * np.sin(pts[:, k])
elif i == j:
vals = -vals * np.sin(pts[:, k])
else:
vals = vals * np.cos(pts[:, k])
d2vals.append(vals)
d2vals = np.moveaxis(np.array(d2vals).reshape((ndim, ndim, -1)), -1, 0)
interp = RBFDInterpolator(
pts,
vals,
dvals,
d2vals,
# extra_degree=2,
# kernel='gaussian',
neighborhood_size=28,
clustering_radius=.05
)
from Peeves.Timer import Timer
# from Peeves import BlockProfiler
with BlockProfiler(name="{} pts:".format(len(interp.grid))):
interp(interp.grid[:25] + .2, deriv_order=0, neighbors=5)
RBFForms
def test_RBFForms(self):
def test_1D(fn, pts):
base_interp = RBFDInterpolator.create_function_interpolation(
pts.reshape((-1, 1)),
lambda p,f=fn:f(p).flatten()
)
#
g = np.linspace(np.min(pts)-.3, np.max(pts)+.3, 50)
cp = plt.CompositePlot(
plt.Plot(g, base_interp(g.reshape(-1, 1)), color='red'),
plt.Plot(g, fn(g), color='green', linestyle='-'),
plt.ScatterPlot(pts, fn(pts))
).show(interactive=False)
dinterp = RBFDInterpolator.create_function_interpolation(
pts,
lambda p,f=fn:f(p).reshape(-1),
lambda p,f=fn.deriv(1):f(p).reshape(-1, 1),
lambda p,f=fn.deriv(2):f(p).reshape(-1, 1, 1)
)
#
itp = dinterp
g = np.linspace(np.min(itp.grid)-.3, np.max(itp.grid)+.3, 50)
cp = plt.CompositePlot(
plt.Plot(g, itp(g.reshape(-1, 1)), color='red'),
plt.Plot(g, fn(g), color='green', linestyle='dashed'),
plt.ScatterPlot(pts, fn(pts))
).show(interactive=False)
sym = Symbols('xyz')
# np.random.seed(3)
# fn = sym.morse(sym.x)
# pts = np.random.uniform(low=-.5, high=1.2, size=30)
# test_1D(sym.morse(sym.x), pts)
# test_1D(sym.sin(sym.x), pts)
# test_1D(sym.morse(sym.x)*sym.sin(sym.x), pts)
#
# raise Exception(...)
np.random.seed(3)
ndim = 2
pts = np.random.uniform(low=-.5, high=1.2, size=(10000, ndim))
np.random.seed(3)
new = np.random.uniform(low=-.6, high=1.3, size=(500, ndim))
# fn = sym.morse(sym.x) * sym.morse(sym.y) - sym.morse(sym.x) - sym.morse(sym.y)
for fn in [
sym.morse(sym.x) * sym.morse(sym.y),
sym.morse(sym.x) * sym.morse(sym.y) - sym.morse(sym.x) - sym.morse(sym.y)
]:
dinterp = RBFDInterpolator.create_function_interpolation(
pts,
fn,
lambda p,f=fn.deriv(order=1):f(p).transpose(),
lambda p,f=fn.deriv(order=2):f(p).transpose(2, 0, 1),
clustering_radius=1e-5
)
# dinterp2 = RBFDInterpolator.create_function_interpolation(
# pts,
# fn,
# lambda p, f=fn.deriv(order=1): f(p).transpose(),
# lambda p, f=fn.deriv(order=2): f(p).transpose(2, 0, 1),
# multicenter_monomials=False
# )
vals = dinterp(new, deriv_order=2)
print_errors = True
val_diff = vals[0] - fn(new)
if print_errors:
print("avg diff:", np.average(val_diff))
print("median diff:", np.median(val_diff))
print("std diff:", np.std(val_diff))
self.assertLess(
np.abs(np.average(val_diff)),
.01,
msg='failed for {}'.format(fn)
)
self.assertLess(
np.std(val_diff),
.05,
msg='failed for {}'.format(fn)
)
grad_diff = vals[1] - fn.deriv(order=1)(new).T
if print_errors:
print("avg grad diff:", np.average(grad_diff))
print("median grad diff:", np.median(grad_diff))
print("std grad diff:", np.std(grad_diff))
# bad_pos = np.where(np.abs(grad_diff) > .5)[0]
# good_pos = np.setdiff1d(np.arange(len(dinterp.grid)), bad_pos)
#
# plt.ListTriPlot3D(*dinterp.grid.T, dinterp.vals).show()
#
# bad_inds = dinterp.get_neighborhood(new[bad_pos[0]:bad_pos[0]+1], neighbors=15)[0]
# plt.ListTriPlot3D(*dinterp.grid[bad_inds].T, dinterp.vals[bad_inds])
#
# good_inds = dinterp.get_neighborhood(new[good_pos[0]:good_pos[0]+1], neighbors=15)[0]
# plt.ListTriPlot3D(*dinterp.grid[good_inds].T, dinterp.vals[good_inds]).show()
# raise Exception(np.where(np.abs(grad_diff) > .5))
self.assertLess(
np.abs(np.average(grad_diff)),
.05,
msg='failed for {}'.format(fn)
)
self.assertLess(
np.std(grad_diff),
.2,
msg='failed for {}'.format(fn)
)
hess_diff = vals[2] - fn.deriv(order=2)(new).transpose(2, 0, 1)
if print_errors:
print("avg hess diff:", np.average(hess_diff))
print("median hess diff:", np.median(hess_diff))
print("std hess diff:", np.std(hess_diff))
self.assertLess(
np.abs(np.average(hess_diff)),
.2,
msg='failed for {}'.format(fn)
)
self.assertLess(
np.std(hess_diff),
.8,
msg='failed for {}'.format(fn)
)
Symbolics
def test_Symbolics(self):
from McUtils.Misc import njit
sym = Symbols('xyz')
x, y, z = sym.vars
e = sym.cos(x)
c = e.compile()
# pts = np.random.rand(10000)
#
# with Timer("base"):
# e(pts)
# with Timer('comp'):
# c(pts)
#
#
# import astunparse
# print(
# astunparse.unparse(e.deriv().get_compile_spec())
# )
# raise Exception('...')
d = e.deriv()
pts = np.array([1, 2, 3])
np.testing.assert_allclose(e(pts), np.cos(pts))
np.testing.assert_allclose(d(pts), -np.sin(pts))
m = sym.morse(x)
np.testing.assert_allclose(m(pts), (1-np.exp(-pts))**2)
np.testing.assert_allclose(m.deriv()(pts), 2*np.exp(-pts)*(1 - np.exp(-pts)))
e = sym.cos(x) + sym.cos(y)
pts = np.array([[1, 0], [2, 1], [3, 2]])
np.testing.assert_allclose(
e(pts),
np.cos(pts[:, 0]) + np.cos(pts[:, 1])
)
d = e.deriv()
np.testing.assert_allclose(
d(pts),
np.array([
-np.sin(pts[:, 0]),
-np.sin(pts[:, 1])
])
)
import sympy
np.random.seed(0)
new_pts = np.random.rand(3, 2)
comp_expr = sym.morse(x)*sym.morse(y)# + sym.morse(x) - sym.cos(y)
sx, sy = sympy.symbols(["x", "y"])
sympy_expr = (1-sympy.exp(-sx))**2 *(1-sympy.exp(-sy))**2
# print(
# comp_expr(new_pts),
# sympy.lambdify([x, y], sympy_expr)(
# new_pts[:, 0],
# new_pts[:, 1]
# )
# )
np.testing.assert_allclose(
comp_expr(new_pts),
sympy.lambdify([sx, sy], sympy_expr)(
new_pts[:, 0],
new_pts[:, 1]
)
)
# print(comp_expr.functions[0].tree_repr())
comp_dexpr = comp_expr.deriv()
np.testing.assert_allclose(
comp_dexpr(new_pts),
np.array([
sympy.lambdify([sx, sy], sympy_expr.diff(sx))(
new_pts[:, 0],
new_pts[:, 1]
),
sympy.lambdify([sx, sy], sympy_expr.diff(sy))(
new_pts[:, 0],
new_pts[:, 1]
)
])
)
new_pts = np.random.rand(1000, 2)
with Timer(tag="Custom", number=25):
for _ in range(25):
dexpr_res = comp_dexpr(new_pts)
comp_dexpr_compiled = comp_dexpr.compile()
# import astunparse
# raise Exception(astunparse.unparse(comp_dexpr.get_compile_spec()))
# compiled_exprs = [
# comp_dexpr.functions[0].compile(),
# comp_dexpr.functions[1].compile()
# ]
# raise Exception(compiled_exprs[0](new_pts))
comp_dexpr_compiled(new_pts) # to precompile
with Timer(tag="Compiled", number=25):
for _ in range(25):
comp_res = comp_dexpr_compiled(new_pts)
# raise Exception(comp_res)
exprs = [
sympy.lambdify([sx, sy], sympy_expr.diff(sx)),
sympy.lambdify([sx, sy], sympy_expr.diff(sy))
]
with Timer(tag="SymPy", number=25):
for _ in range(25):
sympy_res = np.array([
e(
new_pts[:, 0],
new_pts[:, 1]
)
for e in exprs
])
self.assertTrue(
np.allclose(dexpr_res, sympy_res)
)
self.assertTrue(
np.allclose(dexpr_res, comp_res)
)
CoordinateFunctions
def test_CoordinateFunctions(self):
fun = (
CoordinateFunction.morse((0, 1), re=1.8, a=1, de=10)
+ (CoordinateFunction.morse((0, 2), re=1.8, a=1, de=10) / 100)
) * CoordinateFunction.sin((1, 0, 2), n=2)
test_coords = np.random.normal(0, 1, size=(1000, 5, 3))
r, morse_vals = fun(test_coords, reexpress=False, order=2)
# base_fig = plt.ScatterPlot(
# r[0][..., 0],
# morse_vals[0],
# plot_range=[None, [-10, 10]]
# )
# plt.ScatterPlot(
# r[0][..., 0],
# morse_vals[1][..., 0],
# figure=base_fig,
# plot_range=[None, [-10, 10]]
# )
# base_fig = plt.ScatterPlot(
# r[0][..., 0],
# morse_vals[2][..., 0, 0],
# figure=base_fig,
# plot_range=[None, [-10, 10]]
# )
# base_fig.show()
# return
base_fig = plt.ListContourPlot(
np.concatenate([r[0], morse_vals[0][..., np.newaxis]], axis=1),
colorbar=True
)
plt.ScatterPlot(
r[0][..., 0],
r[0][..., 1],
figure=base_fig
).show()
InternalReexpansions
def test_InternalReexpansions(self):
import McUtils.Numputils as nput
nat = 5
coords = np.random.rand(nat, 3)
nx = np.prod(coords.shape, dtype=int)
grad = np.random.rand(nx)
hess = np.random.rand(nx, nx)
hess = hess @ hess.T
sample_coords = [
[0, 1], # bond length
[0, 2],
[0, 3],
[0, 1, 2], # angle between the first three atoms
[0, 1, 3],
[1, 2, 3]
]
vals = []
jacs = []
jac_derivs = []
for idx in sample_coords:
if len(idx) == 2:
fun = nput.dist_vec
elif len(idx) == 3:
fun = nput.angle_vec
elif len(idx) == 4:
fun = nput.angle_vec
else:
raise ValueError("no dispatch")
d0, d1, d2 = fun(coords, *idx, order=2)
vals.append(d0)
jacs.append(d1)
jac_derivs.append(d2)
raise Exception(np.array(jac_derivs).shape)
new_surf = nput.tensor_reexpand(
[np.array(jacs), np.array(jac_derivs)],
[grad, hess],
2
)
raise Exception(new_surf)